A Mobile Based Diet Advisory and Obesity Management System
by Bruno (Working Notes)
miRNA Ontology
by Jaycee (Working Notes)
miRNA Co-Regulation Prediction
by Jiang & Jaycee (Working Notes)
miRNA Target Prediction
by Chunxiao (Literature Review, Features and Data, Working Notes)
Computational Construction of MicroRNA Regulation Network in Obesity and Cancer
by Jiang Shu (Working Notes)
Novel Sequencing Analytical Pipeline for MicroRNA Cross-Species Discovery
by Hanyuan Zhang (Working Notes, Working Notes2)
by Kevin Chiang (Working Notes)
Dietary MicroRNA Databases (DMD): An archive database and analytic tool for food-borne microRNAs
by Kevin Chiang
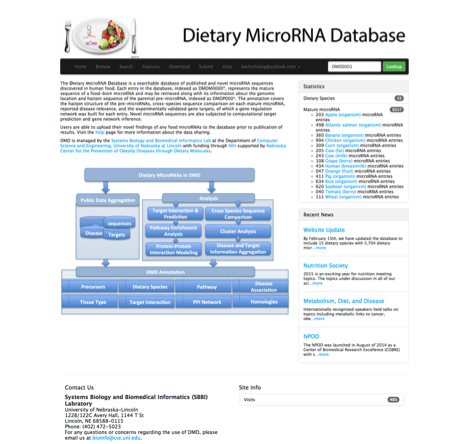 With the advent of high throughput technology, a huge amount of microRNA information has been added to the growing body of knowledge for non-coding RNAs. Here we present the Dietary MicroRNA Databases (DMD), the first repository for archiving and analyzing the published and novel microRNAs discovered in dietary resources. Currently there are fifteen types of dietary species, such as apple, grape, cow milk, and cow fat, included in the database originating from 9 plant and 5 animal species. Annotation for each entry, a mature microRNA indexed as DM0000*, covers information of the mature sequences, genome locations, hairpin structures of parental pre-microRNAs, cross-species sequence comparison, disease relevance, and the experimentally validated gene targets. Furthermore, a few functional analyses including target prediction, pathway enrichment and gene network construction have been integrated into the system, which enable users to generate functional insights through viewing the functional pathways and building protein-protein interaction networks associated with each microRNA. Another unique feature of DMD is that it provides a feature generator where a total of 414 descriptive attributes can be calculated for any given microRNA based on their sequences and structures. DMD would be particularly useful for research groups studying microRNA regulation from a nutrition point of view. The database can be accessed at http://sbbi.unl.edu/dmd/.
With the advent of high throughput technology, a huge amount of microRNA information has been added to the growing body of knowledge for non-coding RNAs. Here we present the Dietary MicroRNA Databases (DMD), the first repository for archiving and analyzing the published and novel microRNAs discovered in dietary resources. Currently there are fifteen types of dietary species, such as apple, grape, cow milk, and cow fat, included in the database originating from 9 plant and 5 animal species. Annotation for each entry, a mature microRNA indexed as DM0000*, covers information of the mature sequences, genome locations, hairpin structures of parental pre-microRNAs, cross-species sequence comparison, disease relevance, and the experimentally validated gene targets. Furthermore, a few functional analyses including target prediction, pathway enrichment and gene network construction have been integrated into the system, which enable users to generate functional insights through viewing the functional pathways and building protein-protein interaction networks associated with each microRNA. Another unique feature of DMD is that it provides a feature generator where a total of 414 descriptive attributes can be calculated for any given microRNA based on their sequences and structures. DMD would be particularly useful for research groups studying microRNA regulation from a nutrition point of view. The database can be accessed at http://sbbi.unl.edu/dmd/. Computational Characterization of Exogenous MicroRNAs That Can Be Transferred into Human Circulation
by Jiang Shu (Notes)
MicroRNAs have been long considered synthesized endogenously until very recent discoveries that human can absorb dietary microRNAs from animal origin. Such compelling evidences, incorporated with controversial discoveries on plant-borne microRNAs, has created high interests in systematic research to study the cross-species transportation of microRNA and explore which and how exogenous microRNAs can get integrated into human circulation and possibly exert functions in humans. Here we present an integrated study where genomics analysis and computational modeling were applied to discover molecular features associated with the mechanism underlying the transportation.
RNA-seq Analytical Pipeline for Gene Expression Summarization and Alternative Splicing Detection
by Jonathan Sherman
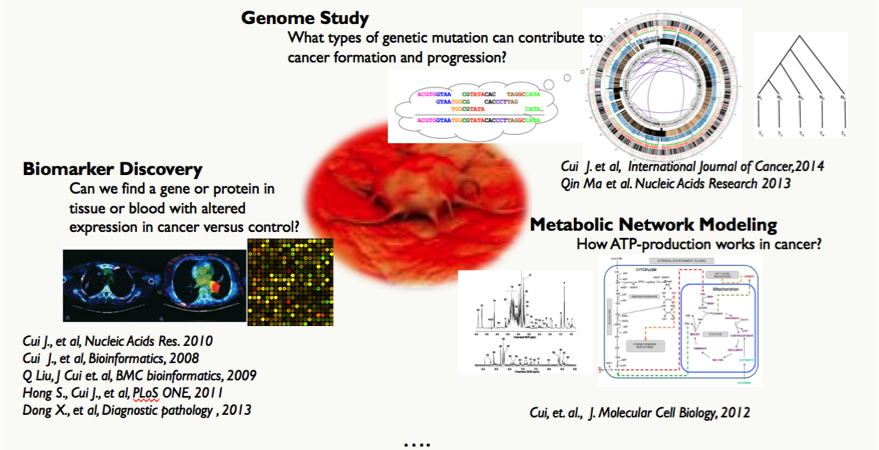
Data-driven Cancer Research
The central goal of our cancer research is to reveal the complex and subtle causal relationships relating environmental factors, genomic predisposition and life style to human cancer formation and progression, through integrated analyses of multiple types of omic and patient data as well as computational modeling.